An app that provides a powerful computational tool to be used for optimising enrollment efficiency in biomarker-driven trials testing the efficacy of targeted therapies has been developed and validated, according to findings presented at the International Congress on Targeted Anticancer Therapies (TAT) in Paris, France, which is organised by the European Society for Medical Oncology (ESMO).
Luca Mazzarella, Drug Development, Experimental Oncology, Istituto Europeo di Oncologia in Milan, Italy and an international team of investigators have proposed a new approach to design biomarker-driven trials evaluating targeted agents that takes into tackling the problem that many of the necessary genetic biomarkers often occur rarely in a population of cancer patients, severely limiting accrual.
The team developed an app as a computational tool, the package Precision Trial Designer (PTD) using the R programming language. They used TCGA data to demonstrate the potential in a simulated 10-arm imaginary trial of multiple cancer types. The trial included genetic alterations that were suggested by the 2017 Molecular Analyses for Personalised Therapy (MAP) Conference and validation as done by comparing the PTD predictions to the actual data compiled in the SHIVA trial.1 The PTD has been deployed as an open access, web-based app using the Shiny-R Studio platform.
The PTD estimates the number of patients needed to screen to recruit and achieve adequate sample size
The app also takes into account an aspect of umbrella trials of targeted therapies, which test multiple hypotheses to allow greater recruitment. While umbrella design increases the trial size, the strategy regarding the order of allocating drugs in the co-existence of multiple actionable mutations, which may affect sample size, should be worked out based on the relative mutation frequencies. As the trial grows in size so does the possibility of conflicting drug allocation, which may alter the statistical power of the trial and yield inconclusive results.
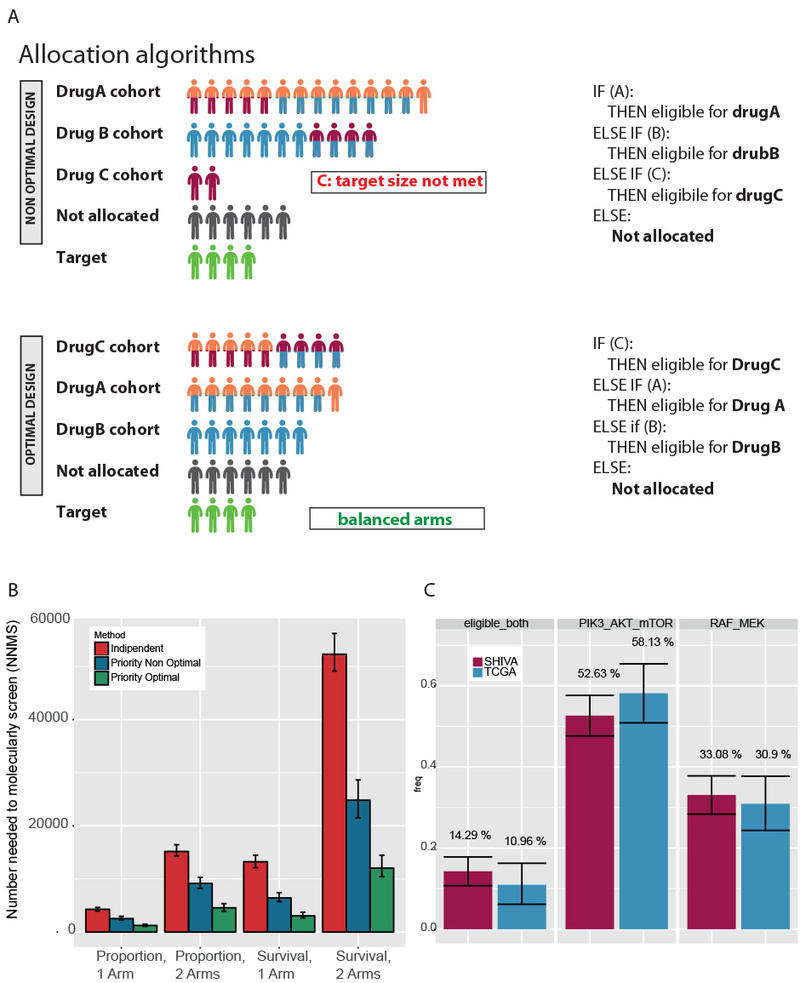
The PTD allows for the estimation of several of the parameters of trial design for evaluating targeted therapies, including the proportion of patients with one or more actionable genetic alteration, the frequency matrix of mutation co-occurrence, and the number of patients needed to molecularly screen (NNMS) for a specific trial design (time-to-event or proportion-based).
The app also provides an allocation rule that maximizes patient accrual by systematically assigning patients to the drug associated with the rarer mutation.
Estimations by the PTD compare favourably with actual data from the SHIVA trial
The investigators found that the PTD optimal design reduced the NNMS by up to 3.5 times (71.8%) in the MAP imaginary trial, as compared with the non-optimal designs. In the SHIVA trial, the PTD correctly predicted the fraction of patients with actionable alteration; the NNMS was 33.51% versus 32.92% by PTD and real SHIVA data (within 95% confidence interval).
The PTD also made correct predictions for the allocation of patients to specific treatments based on the patients' specific genetic alterations.
Conclusions
Based on the findings contained in this poster, the authors concluded that the PTD is able to correctly estimate crucial parameters that allow issues for designing precision oncology trials to be overcome.
They further noted that the availability as an open-access web app provides a useful resource for the entire community of clinical researchers participating in the field of targeted therapy.
Citation
- Le Tourneau et al. Lancet Oncol 2015; 16(13):1324-1334.
Disclosure
This study was funded by the Italian Ministry of Health RF-2013-02357231.
Reference
Mazzarella L, Melloni G, Guida A, et al. Precision trial designer-web: A web-based app to assist in the design of genomics-driven trials. Abstract 118P. International Congress on Targeted Anticancer Therapies (TAT), 5-7 March 2018, Paris, France.